Potential Energy Surfaces
A mildly mathematical introduction to chemistry's coolest hypersurfaces
In part three of my encoding the physical world series, we’ll be tackling potential energy surfaces - from their mathematical construction to their computational applications. Besides their role in modeling molecular dynamics (which I rambled on about here), potential energy surfaces play an indispensable role in statistical mechanical approaches to problems of chemical reactivity, specificity, and conformational stability.
This is a big one, so buckle up :)
Setting the Stage
We start with some quick orienting context - a definition, a justification, a complication, and a comment on perspective.
What it is: a potential energy surface (PES) is a function which maps the potential energy of a molecular system to its geometric configuration across all possible atomic arrangements. Each point on this high dimensional surface represents a specific molecular conformation and its corresponding potential energy.
Why it’s cool: the ability to mathematically model a molecule’s state space with respect to energy turns out to be incredibly useful for mapping and traversing possible reaction pathways. Empirical results can only cover so much ground after all; being able to systematically model the space of possibilities allows us to strategically design experimental work and explore otherwise inaccessible pathways through simulation.
Why it’s hard: high dimensional mathematical representations are often hard to solve and discretize in a way that ensures computational results reflect reality.
Useful meta context: I have a personal skew towards applications in molecular biology and so relevant techniques will be preferentially explored, but the concepts discussed are fundamental to statistical mechanics and applicable across many domains of applied chemistry and physics. My resource pool consequently consists of lectures from materials science, nuclear engineering, and similar alongside those from drug discovery.
Some Prerequisite Theory
Defining Equilibrium
Molecules exist in a state of constant, probabilistic motion. We call the various shapes a molecule takes on in the midst of this motion ‘conformations.’
Though it is tempting to imagine equilibrium as kinetically settled state, you may have some intuition for the conditions under which an ever-moving structure might be said to be in ‘equilibrium’ - its motion appears predictable over some time scale and its state seems to average out to some preferred set of conformations.
But in fact, we can say a little more thanks to the law of detailed balance. The law of detailed balance, a fundamental principle of kinetics, states that in equilibrium the number of occurrences of each reaction in the forward direction is the same as that in the reverse direction. A ‘reaction’ here identifies any sort of molecular interaction such as collision, absorption, emission, or structural change. Detailed balance says that at both the macroscopic and microscopic levels, the system’s reactionary behavior is symmetric with respect to time: the system state would appear the same whether time flows forwards or backwards.
Now, in molecular modeling we often observe molecules in groups; we call these groups ‘ensembles.’ An ensemble of molecules is considered to be in equilibrium when the conformations of the ensemble at any given instant are clustered around a set of energetically favorable states. That is, each low potential energy conformational state observed is populated according to its Boltzmann distribution. The law of detailed balance guarantees this has been achieved mathematically when the probability flux between any two conformational states is zero for all elements of the ensemble.
We will explore later what precisely we mean by ‘low’ potential energy states and how we identify them in the context of all possible potential energy states.
Experimentally, we can use temperature to affect the conformational distributions within an ensemble of molecules - either disturbing or enhancing the equilibrium state depending on the directionality and rate of the temperature change. At higher temperatures, the molecules become more active and the range of accessible conformational states broadens, while at lower temperatures they become less active and the range of accessible conformational states narrows. This is the secret behind powerful imaging techniques like cryo-EM, where rapid vitrification ‘freezes’ molecules in their most probable equilibrium conformations, allowing for comprehensive equilibrium state structural investigations.
Modeling Potential Energy
To understand how we model the potential energy of our system, we need understand all of the different places it lives and what sort of manipulations affect it.
If you’ll recall from our discussion of force fields in Molecular Dynamics, to understand the ways a given molecule can possibly move (and so all the possible conformations it can take on), we need to think about each of the possible variables we can manipulate and the energy required to perform such manipulations:
bond stretching
bond angle bending
bond twisting
Bond Stretching
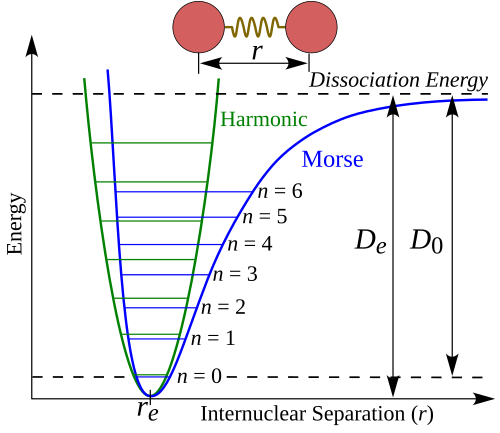
As you’ll recall, bond stretching and angle bending can both be simply approximated with harmonic potentials. However, especially at distances further from equilibrium, bond stretching is more accurately represented by a Morse Potential, a function which models the potential energy between a single pair of atoms with respect to the distance between their nuclei, which can be approximated near its minima by a Taylor expansion of the function.
Given internuclear distance r, equilibrium distance r_e, equilibrium well depth D_e, and well width a, the Morse potential is given by:
The attributes of this potential curve represent important information about the geometry of the underlying structure. The depth of the well is dependent on the bond type (ex: a double bonds yields a shallower well), while the tightness of the well (given by the second derivative) is dependent on the force constant, or the bond stiffness. The minimum point of the well indicates the equilibrium bond distance.
Angle Bending
Angle bending, meanwhile, is generally sufficiently represented by a simple harmonic potential of the form:
Where θ is the angle, θ₀ is the equilibrium angle, and k_θ is the force constant which is given by the angle stiffness.
Bond Torsion
As a rotationally dependent measure, torsion is a periodic function of the dihedral angle derived from the Fourier series of the torsional energy profile. Recall that the dihedral angle here is the angle formed between the planes defined by two sets of sequentially bonded atoms.
The torsional energy of a bond varies as a function of the electron cloud interactions and steric effects that occur during bond rotation. Given bonded molecules A-B-C-D, dihedral angle ω, periodicity n in a 360° rotation, and a barrier height V_n for rotation around the B-C bond, the Fourier series is generally given by:
A phase angle term is often included in the cosine argument as well for greater accuracy.
As with our potential curve, the attributes of our periodic graph tell us important information about the geometry of the underlying structure. For instance, for certain atomic quartets whose rotation is periodic by 120°, that is n=3, the function will reach an anti-minimum and two gauche minima. At a gauche minimum, adjacent substituents sit at a 60° dihedral angle and are staggered similarly to the 180° anti-conformation, but the molecular state is not as stable due to steric strain - the repulsive force of two too-close-together electron clouds. This sort of periodicity is characteristic, for example, of methyl group rotations around a single bond and atom quartets with three fold rotational symmetry around an axis.
Putting It All Together
Parametric Complexity
The maximum number of parameters required to model every possible combination of these manipulations goes up exponentially for n atom types in a given molecule:
the number of bond stretching parameters may go up as much as n^2, if bond stretching is accounted for with every possible pair of atom types
the number of angle bending parameters may go up as much as n^3, if angle bending is accounted for with every possible triplet of atom types
the number of torsion parameters may go up as much as n^4, if torsion is accounted for with every possible quartet of atom types
Modeling Strain
As much as possible, a molecule at equilibrium has zero strain. Strain is a phenomenon that occurs when the individual components of a molecule’s geometry (bond lengths, angles, and torsions) are unable to achieve their ideal values due to constraints imposed by the molecule’s overall geometry. The optimization of one will lead to the deviation of another, and so on. Unable to reach a transferable ideal, the molecule adopts a compromise conformation that distributes strain across all parameters to achieve the lowest possible energy state.
Constructing a Surface
We can now extrapolate the intuition we’ve built from individual attribute curves to a higher dimensional surface representing a molecule’s entire conformational state space. As we move along the continuous surface, we will explore different possible conformations bearing different total potential energies. The deeper the well we end up in, the more stable the structure is likely to be.
Exploring the Surface
Though energetically preferable, molecules do not constantly remain in a state of equilibrium; they must dynamically react to the ever changing conditions of their environment. External conditions such as temperature, pressure, light, and interactions with other molecules can all push molecules out of their equilibrium states. As molecules take on responsive conformations, they express interesting and experimentally relevant behaviors. Understanding the nature of such conformation-dependent behaviors can, for example, help researchers elucidate the mechanistic functions of a given biomolecule.
In a way, the higher order challenge of much of chemistry seems to be: what conditions do/do not cause a particular system to depart from equilibrium, and to what extent? The knowledge of this is of course what fundamentally allows you to understand molecular systems and/or manipulate them towards desired outcomes.
Experimentally, we often consider scenarios where we are moving across an energy surface from the bottom of one well to the bottom of another, which is characteristic of many reaction processes. A feasible reaction pathway can thus be identified along a potential energy surface by finding a saddle point between two wells, specifically a first order saddle point where the surface is at a maximum along exactly one direction (the reaction coordinate) and at a minimum along all other orthogonal directions. At least one such point is guaranteed to exist between any pair of local minima on a continuous energy surface, and is the mathematical definition of a molecular transition state.
Energy Surfaces, Computed
Although helpful to build your intuition for the PES by imagining it as a nice two dimensional surface resting in three dimensions, that’s not actually what’s going on here.
Recall that the dimensionality of the surface can be determined by calculating the molecule’s internal degrees of freedom: for a molecule with N atoms, the total number of dimensions in the potential energy surface is 3N - 6 for non-linear molecules and 3N - 5 for linear molecules.
Due to the high dimensionality of most molecules’ PES, directly calculating all possible configurations and energies is often computationally intractable. Although the transformation from a potential energy function U to a probability distribution P is straightforward in principle:
Where P(x) is the probability of a given configuration x and Z is the partition function:
The problem is calculating Z, which requires integrating over all possible configurations of the system. This is where statistical modeling becomes incredibly useful: we instead approach a solution to this integral by strategically sampling points along the surface.
PES vs. FEL
Briefly, you may have noticed the usage of the thermodynamic beta, β, in the equations above and you’d be wise to point out that a true potential energy surface is defined only with respect to atomic coordinates.
The incorporation of β introduces temperature and entropy as factors in our surface construction. This surface, a Boltzmann-weighted version of our PES, is called a free energy landscape.
Whereas potential energy is stored energy due to position or configuration, free energy is the portion of that energy available to do work at constant temperature and pressure. In the context of understanding biologically relevant processes, free energy differences often play a more significant role than purely potential energy differences. This distinction will become important as we move into sampling methods for simulations - most sample from the FEL rather than the PES.
Markov Chain Monte Carlo
Markov Chain Monte Carlo is a common statistical technique for sampling problems like this, and it’s the one used most often in free energy surface modeling.
The General Sampling Process
Markov Chain Monte Carlo (MCMC) algorithms draw samples from a target probability distribution according to a random variable with probability density proportional to some known function. Each sample is a step in a Markov chain of events whose distribution approximates the target distribution.
Beginning with an initial state selected at random, a new state is proposed according to some proposal distribution. If accepted, a move will be made to that new state, and the process will continue. The decision to accept or deny a new state is made according to some probabilistic acceptance criteria dependent on the state of the current sample. This process is repeated for some specified number of steps; the more steps, the closer the distribution will approximate the target.
Sampling From a Free Energy Landscape
In our case, the target distribution being sampled using MCMC here is a molecule’s FEL and the states being sampled are its various conformations.
The Metropolis-Hastings algorithm is the most popular MCMC algorithm for sampling a free energy landscape. The Metropolis-Hastings algorithm can draw samples from any distribution of probability density P given that we know a calculable function f which is proportional to it. This function f is the one from which we derive our acceptance criteria: the likelihood of accepting a new sample is determined according to the ratio between f calculated at the current and proposed sample values. Because f is proportional to P, we know that this ratio is equal to the ratio between P calculated at the current and proposed sample values:
Once we have evaluated α, our likelihood of acceptance, we generate our random variable u and accept the new sample if u is less than or equal to min(1, α). We then continue on by assigning the next step of the chain to be the new sample if accepted or the current sample if rejected.
In the case of sampling a free energy landscape, we often operate under a certain symmetry assumption which says that for the given proposal distribution, the probability of the current state given new state is the same as the probability of the new state given the current state. Under this symmetry, the final acceptance probability simplifies from the general Metropolis-Hastings acceptance criterion to the Metropolis criterion. Whereas the general final acceptance criterion is given by min(1, α * correction factor) where the correction factor accounts for the probability asymmetry between the orderings of the states, the symmetric final acceptance criterion is simply given by min(1, α).
This practically means that a sample of lower free energy than the previous will be accepted 100% of the time, and a sample of higher free energy than the previous will be accepted with a probability that decreases exponentially with the energy increase. This construction allows the algorithm to occasionally climb energy barriers while still asymptotically sampling from the Boltzmann distribution.
Visualizing in a simplified 3D space, you could imagine that this looks like selected moves on the surface gradually starting to cluster around the bottoms of wells (the lowest free energy states) while moving back upwards towards local maxima and saddle points every so often.
Enhanced Sampling Methods for Molecular Dynamics
Now, not all free energy landscapes are sufficiently ergodic, meaning that a system may not sample the full surface even given sufficient time. For instance, because traditional MCMC algorithms sample according to the Boltzmann distribution without modifying the underlying FEL, they can encounter limitations with sampling relevant but high energy-barrier conformational states (as the probability of their acceptance is near zero). They can also get caught around metastable states - those which are stable but not completely minimized with respect to free energy; these states are local minima but not true equilibrium states. A good example here, if you’ll recall from earlier on, is a molecule with 120° torsional periodicity which will generally bear metastable conformations at its gauche minima.
Because research objectives often call for the simulation of rare states and reaction pathways, enhanced sampling methods are often used in favor of or in conjunction with standard methods. Unlike standard methods, enhanced sampling processes enable ‘read-write’ access, so to speak, on the free energy landscape. These processes leverage features of the FEL to bias samples in favor of certain pathways, in part by strategic rewrites.
Metadynamics
For example, metadynamics is a technique which reconstructs a free energy landscape by progressively raising the energy minima on a collective variable space - a dimensionally reduced version of the surface - to make them less probabilistically favorable, biasing sampling towards regions with higher free energy to ensure the full surface is ultimately explored. This is accomplished technically by applying small repulsive Gaussian biasing potentials at low free energy states explored over the course of the simulation. These potentials take the general form:
Here, s represents a specific point in the collective variable space at which the total accumulated bias potential is evaluated. W is the height of the Gaussian and σ is the width of the Gaussian along each collective variable. The summation over t’<=t indicates that that the total bias is the accumulation of all Gaussians deposited up to the current time t. The current value of the collective variable, s(t), is used to determine the center for the next Gaussian to be added at time t + Δt.
As suggested by their construction, these potentials essentially accumulate to backfill wells in the surface. Over a long enough timescale, this process effectively ends up providing a negative image of the underlying surface.
Umbrella Sampling
Like metadynamics, this technique leverages biasing potentials to enhance sampling of specific regions. However, rather than relying on a series of time-modulated potentials, a single potential is held fixed along a reaction coordinate for the course of a simulation. This biases the sampling algorithm in a particular direction, typically towards critical regions such as transition states and energy barriers. These potentials have the form:
Where k is the force constant, s is the dynamically changing collective variable, and s_0 is the constant collective variable value at which the restraint is applied.
As a fixed-potential approach, this sampling technique is most often run in a series of simulations with different values of s_0 around which the restraint is applied, whose results can be recombined to produce the path’s full free energy profile.
Replica Exchange
I addressed this in detail in my molecular dynamics piece, but it is better contextualized at this point so it is worth a quick re-mention. Replica exchange is a sampling technique that utilizes temperature variance across replicas of a simulated system to enable more complete explorations of the FEL.
As we discussed at the beginning of this article, temperature can affect the conformational distributions within an ensemble of molecules - a higher system temperature makes it more likely that molecules therein will cross energy barriers that would otherwise be prohibitively high.
Rather than running in strict parallel, the system states of replicas held at different temperatures are exchanged occasionally throughout the course of simulation as determined by a Monte Carlo technique, just as in MCMC. This increases the likelihood that uncommon or difficult to mimic state transitions will occur.
Wrapping Up
There’s actually quite a bit more i’d like to say here, but I get the sense that at a certain length these lovely articles feel too formidable for casual consumption. I am having quite a bit of fun diving into this stuff, and cementing my learnings in writing has proven extremely beneficial to refer back to. Along the way, I am hoping to refine my writing voice and tighten up my structure a bit more, but it’s all a process.
As always, I am continually learning, so please reach out to me with any conceptual or logical corrections.
Thanks for reading along!
- Erika :)
Resources
An extremely helpful set of computational chemistry lectures from a series by Professor Chris Cramer at the University of Minnesota:
A lecture on Monte Carlo Techniques as part of a Materials Kinetics series by Professor John Mauro at Penn State:
A presentation on Structural-ensemble probability refinement using Cryo-EM by Pilar Cossio at the Flatiron Institute:
I actually had the pleasure of attending a seminar with a member of the above lab a couple of weeks ago, which was useful inspiration for the writing of this context-building piece.
My final resource for today is the hardcopy book i’m currently reading on Statistical Mechanics by Shang-Keng Ma. You can preview it on Google Books here.